However, this would cause some issues with x-axis (compare ggplot object to plotly ones). Use plotly to plot multiple line or bar in one plot import plotly. Using ggplot::facet_wrap and then ggplotly to make a plotly object. 1 import pandas as pd 2 import numpy as np 3 import matplotlib. 
Then plotting that along with original data but keep the legend only for class = All. So What I can think of are two possibilities:Īdding the whole data (duplicating whole dataframe) and assigning class of All to them. In that case, we could set showlegend to TRUE for those specific plot(s) and set it to FALSE for the rest and also set the legendgroup to trans so we get a unique but also complete legend.Īs I said, here we do not have that special case. Here we do not have a layer with all the ~class entries nor two plots with no intersection in class which their combination also covers all of them. Plotly does not have facet like ggplot2 so it will add legend for each subplot or you can turn it off for some of them. Subplot(nrows = 2, shareX = TRUE, titleY = TRUE) Layout(yaxis = list(title = as.character(.y)), barmode = "stack") To display, set showlegend parameter of Layout object to True. Showlegend = (.y = "2seater"), legendgroup = ~trans) %>% Plotly - Legends, By default, Plotly chart with multiple traces shows legends. Plot_ly(x = ~cyl, y = ~displ, color = ~trans, colors = "Paired", type = "bar", # fill missing levels w/ displ = 0, cyl = first available valueĬomplete(.x, trans, fill = list(displ = 0, cyl = head(.x$cyl, 1))) %>% Follow M-M's suggestion setting showlegend to TRUE for a single group and legendgroup to trans to link the legend entries between subplots.In this example, it would be great to have virginica, versicolor, and setosa listed left to right in the legend (instead of top to bottom). Use tidyr's complete to fill (non-NA) dummy values for the missing factor levels in each class group. Here is an example image of the legend being positioned too low: Moreover, when placing the legend below the plot, it may look better to have legend items listed horizontally (instead of vertically).The following steps are added to the original MWE: Marker = )įig = tools.Another workaround using the tidyverse. I expect that a unique legend name controls more than one trace in the plot. Plotly Express does not support arbitrary subplot capabilities, instead it supports faceting by a given data dimension, and it also supports marginal charts to display distribution information. 
However this is not good, because if the legend is hidden the traces cannot be accessed from it. Plotly Express is the easy-to-use, high-level interface to Plotly, which operates on a variety of types of data and produces easy-to-style figures. Suggested workarounds: Hide some legend names. Since two traces share a name, they should be mapped to a single name in the legend, and when clicking on that name both traces should get hidden. with 'color', if it is within an aesthetic group it's mapped, but when it is outside that group it is fixed and does not appear in the legend). When I'm running the following code, import plotly import numpy as np fig (rows2, cols1) y np.arange (0,10,1) fig.addtrace (go.Scatter (yy,name'name1'), row1,col1) fig.addtrace (go.Scatter. The result is a plot per trace (correct), mapping of legend to both color and symbol (not desired), and repeated trace names in the legend (I would like the trace names to show as unique names).Įxpected behaviour: No automatic mapping, the user can decide whether to map or not (e.g. I'm working with python and plotly, and I'm trying to add a legend to each of my subplots. I have a plot with a legend with two elements: 'name': 'trace1' and 'name':'trace2' but four actual traces, which I divide in two groups by marker symbol. If traces share names (I think) they should be plotted with a unique legend name and not with one name per trace, repeating the names. The problems: mapping of data to the legend seems automatic and cannot be disabled. 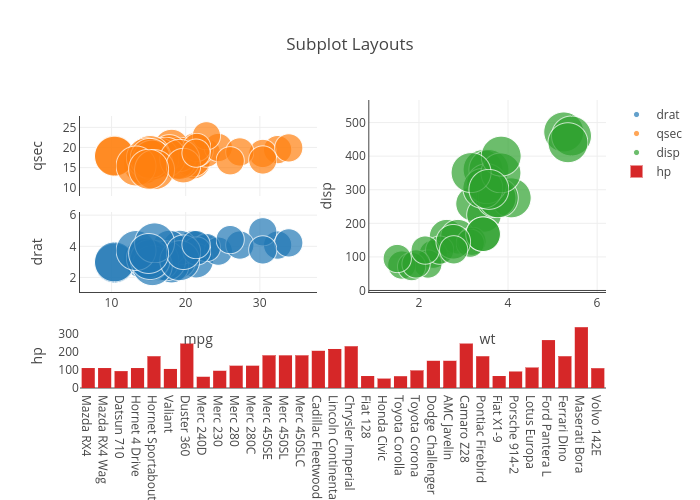
I am not sure whether this is a bug or a feature to request, perhaps it's in between. 
0 Comments
Leave a Reply. |
Details
AuthorWrite something about yourself. No need to be fancy, just an overview. ArchivesCategories |
 RSS Feed
RSS Feed
